● From the very beginning of the proposition of code, it was clear to Francis Crick that there has to be a mechanism to `color{Violet}"read the code"` and also to `color{Violet}"link"` it to the `color{Violet}"amino acids"`, because amino acids have `color{Violet}"no structural specialities"` to read the code uniquely.
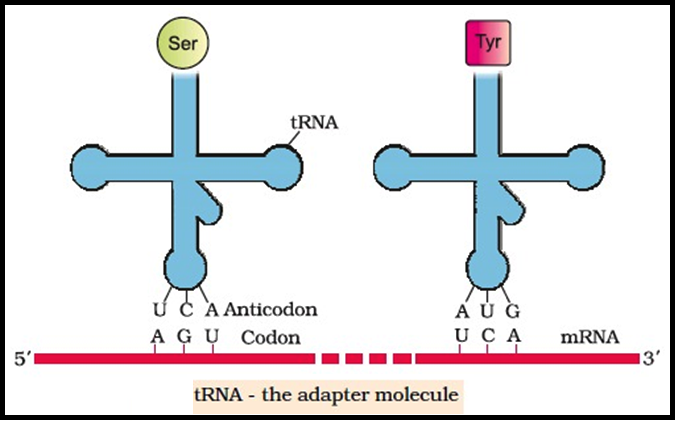
● He postulated the presence of an `color{Violet}"adapter molecule"` that would on one hand read the code and on other hand would bind to specific amino acids.
● The `color{Violet}"tRNA"`, then called `color{Violet}"sRNA"` (`color{Violet}"soluble RNA"`), was known before the genetic code was postulated.
● However, its role as an `color{Violet}"adapter molecule"` was assigned much later.
● tRNA has an `color{Violet}"anticodon loop"` that has bases complementary to the code, and it also has an `color{Violet}"amino acid acceptor"` end to which it binds to amino acids.
● tRNAs are `color{Violet}"specific"` for each amino acid.
● For `color{Violet}"initiation"`, there is another specific tRNA that is referred to as `color{Violet}"initiator tRNA"`.
● There are `color{Violet}"no tRNAs"` for `color{Violet}"stop codons"`.
● In figure, the `color{Violet}"secondary structure"` of tRNA has been depicted that looks like a `color{Violet}"clover-leaf"`.
● In `color{Violet}"actual structure"`, the tRNA is a compact molecule which looks like `color{Violet}"inverted L."`

● From the very beginning of the proposition of code, it was clear to Francis Crick that there has to be a mechanism to `color{Violet}"read the code"` and also to `color{Violet}"link"` it to the `color{Violet}"amino acids"`, because amino acids have `color{Violet}"no structural specialities"` to read the code uniquely.
● He postulated the presence of an `color{Violet}"adapter molecule"` that would on one hand read the code and on other hand would bind to specific amino acids.
● The `color{Violet}"tRNA"`, then called `color{Violet}"sRNA"` (`color{Violet}"soluble RNA"`), was known before the genetic code was postulated.
● However, its role as an `color{Violet}"adapter molecule"` was assigned much later.
● tRNA has an `color{Violet}"anticodon loop"` that has bases complementary to the code, and it also has an `color{Violet}"amino acid acceptor"` end to which it binds to amino acids.
● tRNAs are `color{Violet}"specific"` for each amino acid.
● For `color{Violet}"initiation"`, there is another specific tRNA that is referred to as `color{Violet}"initiator tRNA"`.
● There are `color{Violet}"no tRNAs"` for `color{Violet}"stop codons"`.
● In figure, the `color{Violet}"secondary structure"` of tRNA has been depicted that looks like a `color{Violet}"clover-leaf"`.
● In `color{Violet}"actual structure"`, the tRNA is a compact molecule which looks like `color{Violet}"inverted L."`