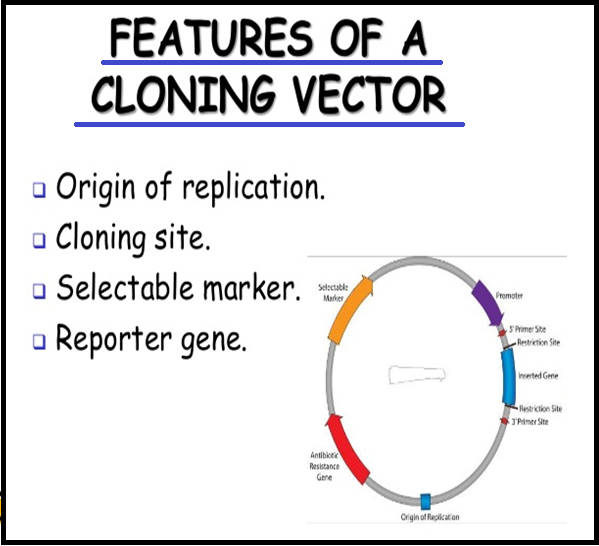
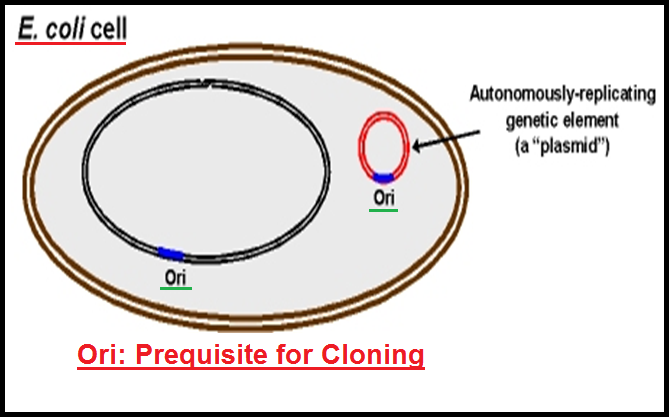
● In order to link the `color{Violet}"alien DNA"`, the vector needs to have very few, `color{Violet}"preferably single, recognition sites"` for the commonly used restriction enzymes.
● Presence of `color{Violet}"more than one"` recognition sites within the vector will generate `color{Violet}"several fragments"`, which will complicate the gene cloning.
● The `color{Violet}"ligation of alien DNA"` is carried out at a `color{Violet}"restriction site"` present in one of the two antibiotic resistance genes.
● For example, you can ligate a `color{Violet}"foreign DNA at the Bam H I"` site of tetracycline resistance gene in the vector `color{Violet}"pBR322"`.
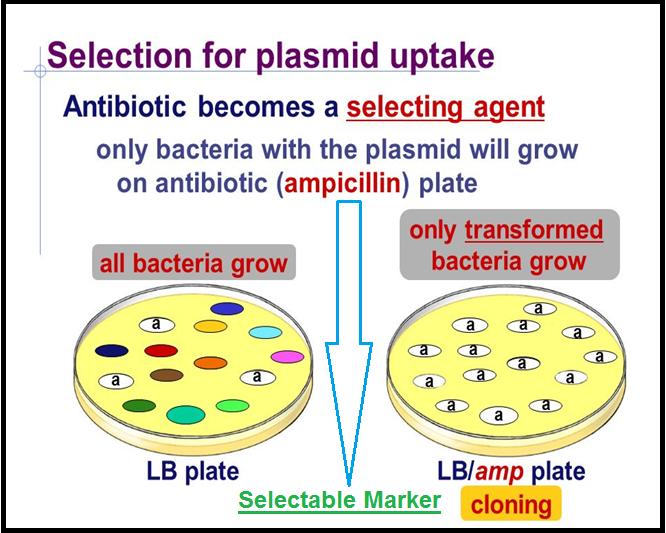
● The `color{Violet}"recombinant plasmids"` will lose `color{Violet}"tetracycline resistance"` due to insertion of foreign DNA but can still be selected out from `color{Violet}"non-recombinant ones"` by plating the transformants on ampicillin containing medium.
● The transformants growing on `color{Violet}"ampicillin"` containing medium are then transferred on a medium containing `color{Violet}"tetracycline"`.
● The recombinants will grow in `color{Violet}"ampicillin containing medium"` but not on that containing tetracycline.
● But, `color{Violet}"nonrecombinants"` will grow on the medium containing `color{Violet}"both the antibiotics"`.
● In this case, one antibiotic resistance gene helps in `color{Violet}"selecting the transformants"`, whereas the other antibiotic resistance `color{Violet}"selection of recombinants"`.
● Selection of recombinants due to `color{Violet}"inactivation of antibiotics"` is a `color{Violet}"cumbersome procedure"` because it requires `color{Violet}"simultaneous plating"` on two plates having different antibiotics.
● Therefore, `color{Violet}"alternative selectable markers"` have been developed which differentiate recombinants from non-recombinants on the basis of their ability to `color{Violet}"produce colour"` in the presence of a chromogenic substrate.
● In this, a recombinant DNA is inserted within the `color{Violet}"coding sequence"` of an enzyme, `color{Violet}"â-galactosidase"`.
● This results into `color{Violet}"inactivation of the enzyme"`, which is referred to as `color{Brown}"insertional inactivation"`.

● The presence of a `color{Violet}"chromogenic substrate"` gives `color{Violet}"blue"` coloured colonies if the plasmid in the bacteria does not have an insert.
● Presence of insert results into `color{Violet}"insertional inactivation"` of the â-galactosidase and the colonies do not produce any colour, these are identified as `color{Violet}"recombinant colonies"`.
● In order to link the `color{Violet}"alien DNA"`, the vector needs to have very few, `color{Violet}"preferably single, recognition sites"` for the commonly used restriction enzymes.
● Presence of `color{Violet}"more than one"` recognition sites within the vector will generate `color{Violet}"several fragments"`, which will complicate the gene cloning.
● The `color{Violet}"ligation of alien DNA"` is carried out at a `color{Violet}"restriction site"` present in one of the two antibiotic resistance genes.
● For example, you can ligate a `color{Violet}"foreign DNA at the Bam H I"` site of tetracycline resistance gene in the vector `color{Violet}"pBR322"`.
● The `color{Violet}"recombinant plasmids"` will lose `color{Violet}"tetracycline resistance"` due to insertion of foreign DNA but can still be selected out from `color{Violet}"non-recombinant ones"` by plating the transformants on ampicillin containing medium.
● The transformants growing on `color{Violet}"ampicillin"` containing medium are then transferred on a medium containing `color{Violet}"tetracycline"`.
● The recombinants will grow in `color{Violet}"ampicillin containing medium"` but not on that containing tetracycline.
● But, `color{Violet}"nonrecombinants"` will grow on the medium containing `color{Violet}"both the antibiotics"`.
● In this case, one antibiotic resistance gene helps in `color{Violet}"selecting the transformants"`, whereas the other antibiotic resistance `color{Violet}"selection of recombinants"`.
● Selection of recombinants due to `color{Violet}"inactivation of antibiotics"` is a `color{Violet}"cumbersome procedure"` because it requires `color{Violet}"simultaneous plating"` on two plates having different antibiotics.
● Therefore, `color{Violet}"alternative selectable markers"` have been developed which differentiate recombinants from non-recombinants on the basis of their ability to `color{Violet}"produce colour"` in the presence of a chromogenic substrate.
● In this, a recombinant DNA is inserted within the `color{Violet}"coding sequence"` of an enzyme, `color{Violet}"â-galactosidase"`.
● This results into `color{Violet}"inactivation of the enzyme"`, which is referred to as `color{Brown}"insertional inactivation"`.

● The presence of a `color{Violet}"chromogenic substrate"` gives `color{Violet}"blue"` coloured colonies if the plasmid in the bacteria does not have an insert.
● Presence of insert results into `color{Violet}"insertional inactivation"` of the â-galactosidase and the colonies do not produce any colour, these are identified as `color{Violet}"recombinant colonies"`.