● SOME IMPORTANT QUESTIONS:
`star` What exactly is `color{Violet}"dominance"`?
`star` Why are some alleles `color{Violet}"dominant and some recessive"`?
● Every gene, contains the information to `color{Violet}"express a particular trait"`.
● In a diploid organism, there are `color{Violet}"two copies of each gene"`, i.e., as a pair of alleles.
● Now, these two alleles need not always be `color{Violet}"identical"`, as in a heterozygote.
● One of them may be different due to some `color{Violet}"changes that it has undergone"` which modifies the information that particular allele contains.
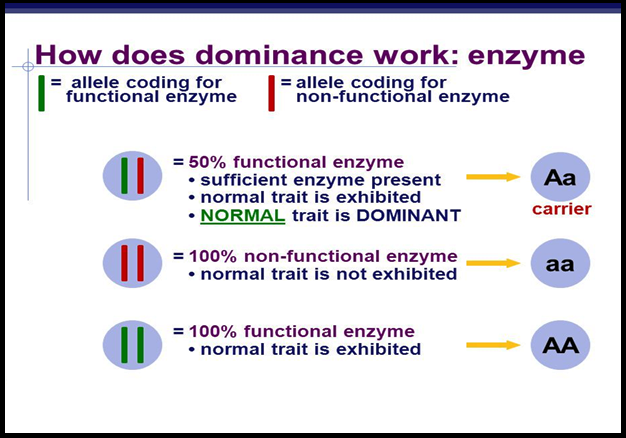
● Let’s take an example of a gene that contains the `color{Violet}"information"` for producing an `color{Violet}"enzyme"`.
● Now there are two copies of this gene, the two `color{Violet}"allelic forms"`.
● Let us assume (as is more common) that the `color{Violet}"normal allele"` produces the `color{Violet}"normal enzyme"` that is needed for the transformation of a `color{Violet}"substrate S"`.
● Theoretically, the `color{Violet}"modified allele"` could be responsible for production of –
(i) the `color{Violet}"normal/less efficient"` enzyme, or
(ii) a `color{Violet}"non-functional enzyme"`, or
(iii) `color{Violet}"no enzyme"` at all
● In the first case, the modified allele is equivalent to the unmodified allele, i.e., it will produce the `color{Violet}"same phenotype/trait"`, i.e., result in the transformation of substrate S. Such equivalent allele pairs are very common.
● But, if the allele produces a `color{Violet}"non-functional enzyme"` or no enzyme, the phenotype may be affected.
● The phenotype/trait will only be dependent on the functioning of the `color{Violet}"unmodified allele"`.
● The unmodified (functioning) allele, which represents the `color{Violet}"original phenotype"` is the dominant allele and the modified allele is generally the recessive allele.
● Hence, in the example above the recessive trait is seen due to `color{Violet}"non-functional enzyme"` or because no enzyme is produced.
● SOME IMPORTANT QUESTIONS:
`star` What exactly is `color{Violet}"dominance"`?
`star` Why are some alleles `color{Violet}"dominant and some recessive"`?
● Every gene, contains the information to `color{Violet}"express a particular trait"`.
● In a diploid organism, there are `color{Violet}"two copies of each gene"`, i.e., as a pair of alleles.
● Now, these two alleles need not always be `color{Violet}"identical"`, as in a heterozygote.
● One of them may be different due to some `color{Violet}"changes that it has undergone"` which modifies the information that particular allele contains.
● Let’s take an example of a gene that contains the `color{Violet}"information"` for producing an `color{Violet}"enzyme"`.
● Now there are two copies of this gene, the two `color{Violet}"allelic forms"`.
● Let us assume (as is more common) that the `color{Violet}"normal allele"` produces the `color{Violet}"normal enzyme"` that is needed for the transformation of a `color{Violet}"substrate S"`.
● Theoretically, the `color{Violet}"modified allele"` could be responsible for production of –
(i) the `color{Violet}"normal/less efficient"` enzyme, or
(ii) a `color{Violet}"non-functional enzyme"`, or
(iii) `color{Violet}"no enzyme"` at all
● In the first case, the modified allele is equivalent to the unmodified allele, i.e., it will produce the `color{Violet}"same phenotype/trait"`, i.e., result in the transformation of substrate S. Such equivalent allele pairs are very common.
● But, if the allele produces a `color{Violet}"non-functional enzyme"` or no enzyme, the phenotype may be affected.
● The phenotype/trait will only be dependent on the functioning of the `color{Violet}"unmodified allele"`.
● The unmodified (functioning) allele, which represents the `color{Violet}"original phenotype"` is the dominant allele and the modified allele is generally the recessive allele.
● Hence, in the example above the recessive trait is seen due to `color{Violet}"non-functional enzyme"` or because no enzyme is produced.